This paper is available on arxiv under CC 4.0 license.
Authors:
(1) Andrea Roncoli, Department of Computer, Science (University of Pisa);
(2) Aleksandra Ciprijanovi´c´, Computational Science and AI Directorate (Fermi National Accelerator Laboratory) and Department of Astronomy and Astrophysics (University of Chicago);
(3) Maggie Voetberg, Computational Science and AI Directorate, (Fermi National Accelerator Laboratory);
(4) Francisco Villaescusa-Navarro, Center for Computational Astrophysics (Flatiron Institute);
(5) Brian Nord, Computational Science and AI Directorate, Fermi National Accelerator Laboratory, Department of Astronomy and Astrophysics (University of Chicago) and Kavli Institute for Cosmological Physics (University of Chicago).
Table of Links
Acknowledgments and Disclosure of Funding, and References
3 Results
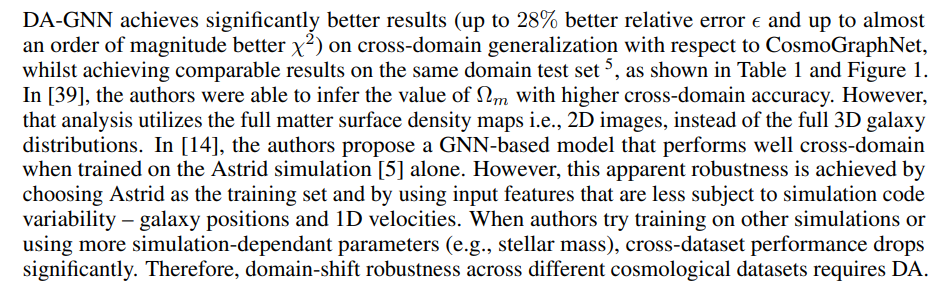

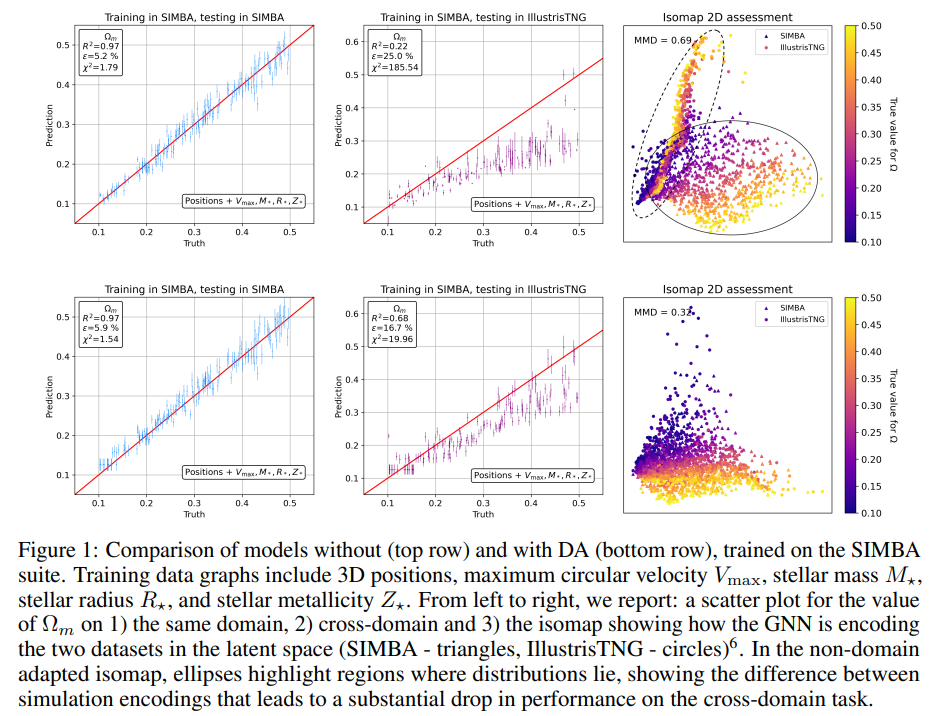
Latent space organization Isomaps are two-dimensional projections of the multi-dimensional latent space [35]. Figure 1 shows the difference in the latent space structure without (top row) and with (bottom row) DA. Ellipses in the top right isomap highlight how the two distributions are encoded in different regions of the latent space. Without the MMD loss, the model encodes samples with very different values of Ωm close to each other, if they originate from different simulations (circles and triangles of different colors are overlapping). This scenario leads to the fragility of the regressor, which cannot learn to output different values for the same latent space encodings. On the contrary, the DA-GNN (bottom right plot) correctly encodes the samples in a domain-invariant way. Visually, circle and triangle distributions are overlapping, which indicates domain mixing. Furthermore, the direction in the color gradient shows that the DA-GNN encodes information such that the regressor can now more correctly predict cosmological parameters based on the encodings of both simulations.
[5] In [41], authors get slightly better results for the same domain, and slightly worse for the cross-domain tests. We impute these differences to choices such as batch sizes and optimization techniques we took due to computational and time constraints.
[6] In Appendix A, the IllustrisTNG counterpart of this plot is presented.
